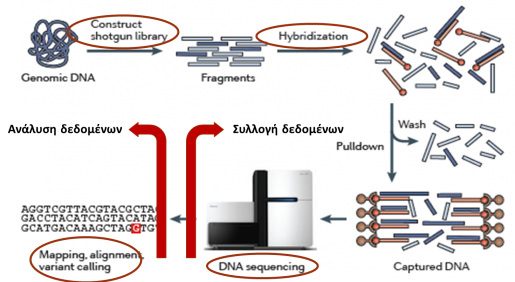
We perform whole exome sequencing (WES), via Next Generation Sequencing (NGS) on a Genome Analyzer – Ion Proton platform.
As a first step, the patient’s DNA is subjected to standard QC procedures, in terms of size distribution and exact DNA quantity. The applied methodology aims at analyzing the DNA sequence of the exons of all known human genes (CCDS genes), as represented in the Ion Ampliseq Exome Sequencing reagent.
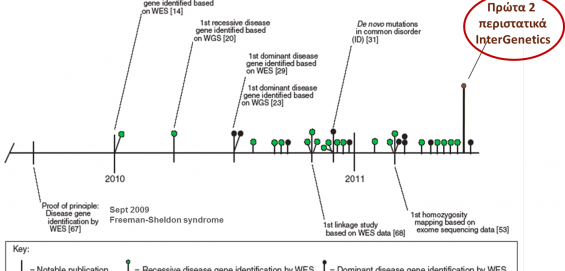
Bearing in mind that the vast majority of pathogenic mutations (>80%) are located in the coding regions – exons of the genes and the exon-intron boundaries, as opposed to deep intronic mutations which are typically non-pathogenic, the technique of whole exome sequencing (WES) permits the simultaneous identification of pathogenic mutations in almost all known human genes related to genetic diseases.
However, it must be noted that the technique, as applied today, is in a position to analyze successfully the DNA sequence of approximately 90-95% of all exons (CCDS), as compared to the Reference Genome (NCBI hg19).
Furthermore, the depth of read and coverage is not the same across all regions, and therefore the confidence of variant calling may vary.
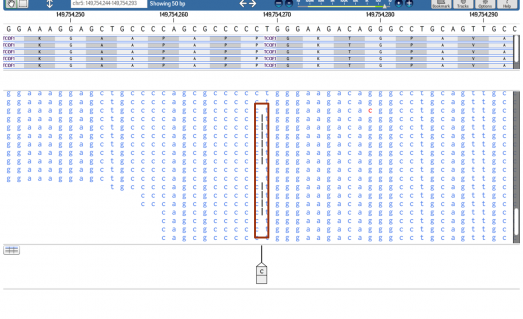
DNA sequence alignment and variant calling is performed with the aid of the Ion Torrent Server Suite, Ion Reporter, Softgenetics NextGene v2.1 and DNAnexus analysis tools.
All genomic variant data are imported, filtered and prioritized using an in-house proprietary Exome Analysis Application (EMA®) software pipeline and novel mutations are assessed for pathogenicity through the pipeline, utilizing a combination of several in silico prediction tools, with reference to numerous public databases and resources and an internal Greek variant frequency database of 120 individuals.
Finally, the test generally evaluates and reports:
- known pathogenic mutations, previously reported in affected individuals and which have been directly linked to the expression of the disease, or
- novel obligatory pathogenic variants/mutations, such as nonsense, frameshift, canonical splice-site variants, indels.
In certain cases, it is possible that other type of variants, not included in any of the above categories, may also be evaluated and reported.
The applied methodology has certain important limitations:
- It may not detect variants in ‘blind spots’ of exons, which may have not been analyzed in the sample (typically <10%) or may be generally refractory to this type of analysis (e.g. highly GC-rich regions, etc.).
- It is not able to reliably detect extensive genomic deletions/duplications (copy number variants).
- It is not in a position to detect triplet-repeat expansions, associated with a certain type of genetic disorders ( triplet-repeat disorders).
- The evaluation and reporting adheres in general to the ACMG guidelines (Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405-24).